Categories
Antibodies
A comprehensive range of monoclonal, polyclonal and secondary antibodies
Kits & Assays
For the study of apoptosis, DNA damage, oxidative stress, angiogenesis, enzyme activity an...
Proteins & Peptides
An extensive range of high quality human and animal protein products
Reagents & Labware
A comprehensive range of life science research reagents and laboratory equipment
What is Epigenetics?
Epigenetics is the study of cellular and physiological phenotypic trait variations that result from external or environmental factors that switch genes on and off and affect how cells express genes. Epigenetic research seeks to describe dynamic alterations in the transcriptional potential of a cell.
Mechanisms that produce such changes are DNA methylation and histone modification, each of which alters how genes are expressed without altering the underlying DNA sequence. Gene expression can be controlled through the action of repressor proteins that attach to silencer regions of the DNA. These epigenetic changes may last through cell divisions for the duration of the cell's life, and may also last for multiple generations even though they do not involve changes in the underlying DNA sequence of the organism; instead, non-genetic factors cause the organism's genes to behave differently.
Epigenetics is important for cellular differentiation, embryology, the regulation of gene expression, aging, cancer, and other diseases.
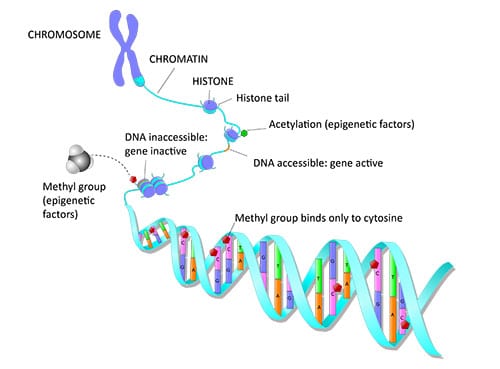
Why Study Epigenetics?
Epigenetics is critical in normal development. All of our cells have the same DNA, yet can be so different from each other. Think of the difference between a muscle cell and a red blood cell for example. How can this be when they contain the same DNA? Cells become specialized during development based on which genes are turned off or turned on resulting in differential gene expression, and ultimately different cellular phenotypes. Epigenetic changes can influence gene expression, primarily at the transcription level.
X chromosome inactivation in female mammals is another critical epigenetic mechanism involved in development, ensuring equal gene dosage between males and females.
As well as the importance in normal processes, epigenetics has been linked to a number of human disorders and diseases. Uncovering and understanding the specific causes of these diseases could lead to revolutionizing treatments or prevention of inherited diseases without altering DNA sequences.
How Can We Study Epigenetics?
Epigenetics is an exciting area of research, and one which is showing rapid advancement. What was once a vague and not well understood area, has become a key area of research, with scientists uncovering the molecular mechanisms underlying epigenetics and working to drive research forward.
Many methods exist to study epigenetics, including assays to assess chromatin configuration and analyse epigenetic modulating enzymes. Chromatin immunoprecipitation (ChIP) assay is an essential tool in epigenetic research which allows for the study of specific protein-DNA interactions.
Our Products
DNA Methylation
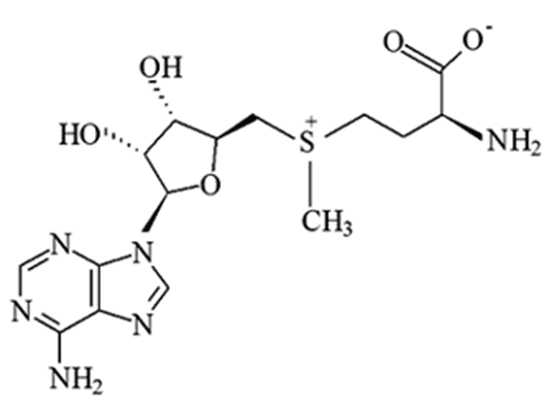
DNA methylation is a key epigenetic mechanism that involves addition of a methyl group to the cytosine of a CpG dinucleotide in DNA. When located in a gene promoter, DNA methylation typically acts to repress gene transcription. Methylation of DNA is required for proper gene regulation during development and is involved in X-chromosome inactivation and allele-specific silencing of imprinted genes. It also plays a crucial role in the development of nearly all types of cancer.
Histone Methylation
Histone methylation is a process by which methyl groups are transferred to amino acids of histone proteins. Methylation of histones can either increase or decrease transcription of genes, depending on which amino acids in the histones are methylated, and how many methyl groups are attached. Methylation events that weaken chemical attractions between histone tails and DNA increase transcription. This is because they enable the DNA to uncoil from nucleosomes so that transcription factors and RNA polymerase can access the DNA. This process is critical for the regulation of gene expression that allows different cells to express different genes.
Histone Acetylation
Acetylation and deacetylation of the lysine residues within the N-terminal tail protruding from the histone core of the nucleosome are essential processes of gene regulation. These reactions are typically catalysed by enzymes with histone acetyltransferase (HAT) or histone deacetylase (HDAC) activity.
Acetylated histones represent a type of epigenetic marker within chromatin. Acetylation removes the positive charge on the histones, thereby decreasing the interaction of the N termini of histones with the negatively charged phosphate groups of DNA. As a consequence, the condensed chromatin is transformed into a more relaxed structure that is associated with greater levels of gene transcription. This relaxation can be reversed by HDAC activity. Relaxed, transcriptionally active DNA is referred to as euchromatin. More condensed (tightly packed) DNA is referred to as heterochromatin.
Bromodomains
Bromodomains are protein domains, found in many proteins associated with transcription and chromatin that recognise acetylated lysine residues. Bromodomains, as the “readers” of lysine acetylation, are responsible in transducing the signal carried by acetylated lysine residues and translating it into various normal or abnormal phenotypes. This recognition is often a prerequisite for protein-histone association and chromatin remodelling, making them important in epigenetics research.